Molecular Descriptors Family on Structure Activity Relationships
4. Molar Refraction of Cyclic Organophosphorus Compounds
Lorentz JÄNTSCHI and Sorana BOLBOACĂ
Technical University of Cluj-Napoca, Romania, http://lori.academicdirect.org
“Iuliu Haţieganu” University of Medicine and Pharmacy, http://sorana.academicdirect.ro
Abstract
The molecular descriptors family on structure activity relationships methodology was applied on ten cyclic organophosphorus compounds in order to predict theirs molar refraction. A number of 107692 significantly different MDF members enter into a multiple linear regression analysis. A pair of descriptors (lGDmSMt, lAmrfEt), which have the best performing ability in prediction of molar refraction of cyclic organophosphorus compounds, was found and a bi-varied MDF SAR model was built. After performing leave-one-out cross-validation, satisfactory result was obtained with cross-validation r2cv and r2 values of 0.9999 and 0.9999. The external validation of the bi-varied MDF SAR model and its ability in prediction of molar refraction of cyclic organophosphorus compounds is demonstrated by the results obtained in training vs. test experiment. The correlated correlation results proved us that the ability in prediction of molar refraction of cyclic organophosphorus compounds with bi-varied MDF SAR model is significantly better compared with the previous reported SAR (see pZ = 0.0 % from Steiger’s Z test). The results showed clearly that the molar refraction of cyclic organophosphorus compounds is almost of topological nature (99.99%), and is strongly dependent on atomic relative mass and atomic electronegativity.
Keywords
Molecular Descriptors Family on Structure Activity Relationships, Molar refraction, Cyclic organophosphorus compounds
Introduction
In chemistry, the molar refraction is an approximate measure of the total volume (without free space) of the molecules in one mole of the compound. The molar refraction is calculated by the Lorenz-Lorentz formula [1]:
((n2-1)/(n2+2))·(M/d)
where n is index of refraction (the ratio between the speed of light in a vacuum and the speed of light through the compound), M is the compound's molecular weight (the weight of one molecule, in atomic mass units), and d is the compound's density (in grams per cubic centimeter).
The molar refraction of ten cyclic organophosphorus compounds was previous studied using the Szeged topological indices [2]. The best reported SAR model has the following characteristics:
· Number of compounds: 10 and number of components: 2 (SZp = a topological descriptor, SZeX = an electronic descriptor);
· SAR model: Ŷ = 36.821 + 0.016·SZp - 0.631·SZeX (Ŷ = estimated molar refraction) and the correlation coefficient: r = 0.9755.
The aim of the research was to test the ability of MDF SAR methodology in prediction of the molar refraction of cyclic organophosphorus compounds and to compare the best performing MDF SAR model with the previous reported SAR model.
Material and Method
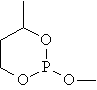
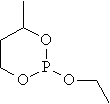
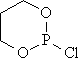
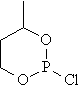
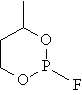
Ten cyclic organophosphorus compounds were studied [2]. The planar structure of the cyclic organophosphorus compounds, molar refraction (MR) and previous estimated molar refraction are in table 1.
Table 1. Planar structure of cyclic organophosphorus compounds, MR and previous estimated molar refraction
|
Mol. |
Cyclic organophosphorus planar structure |
MR |
Estimated MR from ref. [2] |
|
mr01 |
|
35.808 |
36.557 |
|
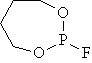
mr02 |
|
40.524 |
40.530 |
|
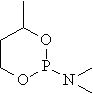
mr03 |
|
30.030 |
32.461 |
|
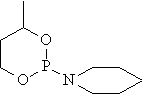
mr04 |
|
34.911 |
33.776 |
|
mr05 |
|
29.222 |
29.896 |
|
mr06 |
|
31.636 |
30.056 |
|
mr07 |
|
43.005 |
40.608 |
|
mr08 |
|
52.029 |
52.084 |
|
mr09 |
|
49.971 |
53.864 |
|
mr10 |
|
58.323 |
54.714 |
The steps of molecular descriptors family on structure activity relationships (MDF SAD) modeling of molar refraction of cyclic organophosphorus compounds were [3]:
· Step 1: Sketch of the cyclic organophosphorus compounds;
· Step 2: Create the cyclic organophosphorus compounds molar refraction file;
· Step 3: Generate cyclic organophosphorus compounds MDF members;
· Step 4: Find the molar refraction SAR models;
· Step 5: Validate the MDF SAR models and compare with previous reported results;
· Step 6: Analyze the selected MDF SAR model.
Results
From a total number of 787968 calculated descriptors, 349553 had real and distinct values. Using a 10-9 significance selector to bias the values, the MDF members were reduced to a number of 107692 significantly different descriptors. In order to obtain the best performing bi-varied MDF SAR model, pairs of MDF members were included into multiple linear regression experiment. The pair of descriptors, which provide best performing prediction of molar refraction of cyclic organophosphorus compounds, contain lGDmSMt and lAmrfEt members. The resulted multiple linear regressions models are available at address:
http://vl.academicdirect.org/molecular_topology/mdf_findings/
A program was developed (j_mdf_demo.php) in order to demonstrate the validity and the complexity of the MDF SAR methodology in prediction of molar refraction of cyclic organophosphorus compounds. The calculations for the first considered cyclic organophosphorus compound (mr01) with lGDmSMt MDF member and for the last considered cyclic organophosphorus compound (mr10) with lAmrfEt member are in Appendix.
The best performing bi-varied MDF SAR model, the calculated valued for the lGDmSMt and the lAmrfEt members, and estimated molar refraction are in table 2. The plot of the bi-varied MDF SAR model from table 2 is in figure 1.
Table 2. The calculated values of the lGDmSMt and lAmrfEt descriptors and estimated by MDF SAR molar refraction
|
Ŷ= 17.39 + 28.25·lGDmSMt – 83.96·lAmrfEt |
|||
|
[ |
|||
|
Mol |
lGDmSMt |
lAmrfEt |
Estimated MR, Ŷ |
|
mr01 |
4.2161 |
1.1978 |
35.912 |
|
mr02 |
4.0257 |
1.0788 |
40.526 |
|
mr03 |
4.7052 |
1.4330 |
29.979 |
|
mr04 |
4.4888 |
1.3004 |
35.001 |
|
mr05 |
4.3624 |
1.3271 |
29.188 |
|
mr06 |
4.4222 |
1.3185 |
31.600 |
|
mr07 |
4.0972 |
1.0740 |
42.949 |
|
mr08 |
3.8446 |
0.8820 |
51.939 |
|
mr09 |
3.9102 |
0.9270 |
50.028 |
|
mr10 |
3.7941 |
0.7890 |
58.336 |
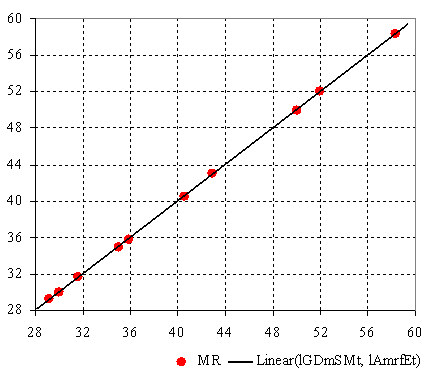
Figure 1. Molar refraction vs. bi-varied MDF SAR estimated molar refraction
The training vs. test experiment was performed in order to validate the bi-varied MDF SAR model. The cyclic organophosphorus compounds were successively split into training and test sets. The experiment run twice times for training sample sizes equal with four, five, six and seven. Corresponding to the training tests sample sizes, the test sets sample sizes were respectively equal with six, five, four, and three. The Multiple Linear Regression results of the experiment are in table 3.
Table 3. Training vs. test sets results using bi-varied MDF SAR model
|
No. |
Training set |
Test set |
||||
|
Molecules |
Intercept |
lGDmSMt |
lAmrfEt |
(1-r2) |
(1-r2) |
|
|
1 |
10, 4, 9, 7 |
19.411 |
27.516 |
-83.055 |
2.5·10-5 |
1.5·10-4 |
|
2 |
9, 6, 5, 7 |
16.578 |
28.473 |
-84.066 |
1.5·10-5 |
4.3·10-5 |
|
3 |
6, 2, 1, 8, 3 |
15.465 |
29.079 |
-85.31 |
3.1·10-5 |
6.2·10-5 |
|
4 |
1, 9, 3, 7, 10 |
15.707 |
28.853 |
-84.739 |
2.2·10-5 |
1.0·10-4 |
|
5 |
8, 1, 9, 4, 5, 6 |
18.617 |
27.821 |
-83.489 |
5.8·10-5 |
2.4·10-5 |
|
6 |
7, 8, 2, 3, 10, 5 |
17.293 |
28.287 |
-83.993 |
1.0·10-5 |
5.7·10-5 |
|
7 |
3, 4, 6, 5, 2, 8, 1 |
17.153 |
28.386 |
-84.258 |
7.9·10-5 |
3.6·10-5 |
|
8 |
10, 9, 4, 7, 3, 6, 5 |
17.908 |
28.024 |
-83.591 |
2.2·10-5 |
6.4·10-6 |
The MDF SAR model (Estimated MR, Ŷ from table 2) has the following associated statistics:
r = 0.9999, r2 = 0.9999, r2adj = 0.9999, r2cv = 0.9999, F = 83012, pF = 4.8·10-14 %,
r2(MR, lGDmSMt) = 0.864, r2(MR, lAmrfEt) = 0.969, r2(lGDmSMt, lAmrfEt) = 0.948
where r is the correlation coefficient, r2 is the squared correlation coefficient, r2adj is the adjusted r2, r2cv is the cross-validation leave-one-out squared correlation score, F is the Fissssher parameter, and pF is the Fisrer’s associated p-value.
A correlated correlations analysis using the Steiger’s Z test was applied in order to compare the bi-varied MDF SAR model with previous reported SAR model. The correlated correlations results are:
r(MR, MDF SAR) = 0.99998; r(MR, SAR) = 0.97537; r(MDF SAR, SAR) = 0.97555;
Z = 9.25276; pZ = 0.0 %
where Z is the Steiger’s Z test parameter and pZ is the associated p-value.
Discussions
The pair of MDF members that have the best ability in prediction of the molar refraction of cyclic organophosphorus compounds contain lGDmSMt and lAmrfEt members. Both members consider the topological shape of the molecules (t), one descriptor takes into consideration atomic relative mass (M) of the molecules while the other takes into consideration atomic electronegativity (E). The squared correlation coefficients computed between molar refraction and MDF members showed that 86% in variation of molar refraction is explainable by its linear relation with lGDmSMt member and almost 97% in variation of molar refraction is explainable by its linear relation with lAmrfEt member. Ninety-nine percent in variation of molar refraction is explainable by its linear relation with the pair (lGDmSMt, lAmrfEt) of MDF members. The probability of a wrong MDF SAR model is equal with 4.8·10-14 %.
The cross-validation leave-one-out correlation score of bi-varied MDF SAR model demonstrate the power of the model in molar refraction of cyclic organophosphorus compounds prediction (r2cv = 0. 9999).
The external validation of the bi-varied MDF SAR model and its ability in prediction of molar refraction of cyclic organophosphorus compounds is demonstrated by the results of training vs. test experiment (table 3). The averages of squared correlation coefficients from training (0.9999) and test (0.9999) are equal, which prove its ability in prediction.
The correlated correlation experiment proved us that the ability in prediction of molar refraction of cyclic organophosphorus compounds with bi-varied MDF SAR model is significantly better compared with the previous reported SAR (see pZ = 0.0 % from Steiger’s Z test).
Conclusions
The molar refraction of cyclic organophosphorus compounds is almost of topological nature (99.99%), and is strongly dependent on atomic relative mass and atomic electronegativity.
Even if the MDF SAR methodology is complex and time consuming the result worth the effort because the ability in prediction of the molar refraction of cyclic organophosphorus compounds with bi-varied MDF SAR model is significantly better comparing with previous reported SAR model.
References
[1] Vakili-Nezhaad G. R., Modarress H., A New Characterization Factor for Hydrocarbons and Petroleum Fluids Fractions, Oil & Gas Science and Technology – Rev. IFP, Vol. 57, No. 2, 2002, p. 149-154.
[2] Jäntschi L., Mureşan S., Diudea M.V., Modeling molar refraction and chromatographic retention by Szeged Indices, Studia Universitatis Babeş-Bolyai, Chemia, 2000, XLV(1-2), p. 313-318.
[3] Jäntschi L., Molecular Descriptors Family on Structure Activity Relationships 1. The review of Methodology, Leonardo Electronic Journal of Practices and Technologies, AcademicDirect, 2005, Issue 6, p. 76-98.
Appendix
The appendix are from output of j_mdf_demo.php program.
Demonstrative calculations for mr01 molecule and lGDmSMt MDF member
POST data: Array
(
[hin] => 01_mr1001.hin
[Do] => t
[Ap] => M
[Df] => S
[Im] => m
[Fc] => D
[Sf] => G
[Lo] => l
)
Molecule's data: m_c Object
(
[a] => 9
[b] => 9
[atom] => Array
(
[1] => Array
(
[0] => 1
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.1095657
[6] => 0.4318282
[7] => 0.6380987
[8] => 2.849884
[9] => 1
[10] => 2
[11] => 1
)
[2] => Array
(
[0] => 2
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => 0.5250869
[6] => -0.2941397
[7] => 0.1247597
[8] => 1.592484
[9] => 3
[10] => 1
[11] => 1
[12] => 3
[13] => 1
[14] => 7
[15] => 1
)
[3] => Array
(
[0] => 3
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.0686326
[6] => -0.2920918
[7] => -1.415741
[8] => 1.596032
[9] => 2
[10] => 2
[11] => 1
[12] => 4
[13] => 1
)
[4] => Array
(
[0] => 4
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => 0.4661818
[6] => -1.039482
[7] => -1.962821
[8] => 0.3649576
[9] => 2
[10] => 3
[11] => 1
[12] => 5
[13] => 1
)
[5] => Array
(
[0] => 5
[1] => -
[2] => O
[3] => O2
[4] => -
[5] => -0.9945602
[6] => -0.3896529
[7] => -1.532219
[8] => -0.8337278
[9] => 2
[10] => 4
[11] => 1
[12] => 6
[13] => 1
)
[6] => Array
(
[0] => 6
[1] => -
[2] => P
[3] => P5
[4] => -
[5] => 1.883921
[6] => -0.5882888
[7] => 0.2352459
[8] => -1.032513
[9] => 3
[10] => 5
[11] => 1
[12] => 7
[13] => 1
[14] => 8
[15] => 1
)
[7] => Array
(
[0] => 7
[1] => -
[2] => O
[3] => O2
[4] => -
[5] => -1.095338
[6] => 0.3748803
[7] => 0.6139225
[8] => 0.4270911
[9] => 2
[10] => 6
[11] => 1
[12] => 2
[13] => 1
)
[8] => Array
(
[0] => 8
[1] => -
[2] => O
[3] => O2
[4] => -
[5] => -1.019472
[6] => 0.6156649
[7] => 1.142236
[8] => -1.997664
[9] => 2
[10] => 6
[11] => 1
[12] => 9
[13] => 1
)
[9] => Array
(
[0] => 9
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => 0.4202826
[6] => 0.2612277
[7] => 2.5274
[8] => -2.022074
[9] => 1
[10] => 8
[11] => 1
)
)
[prop] => Array
(
[1] => Array
(
[0] => 1
[1] => 4
[2] => 12
[3] => 2.746
[4] => 2.6296707827796
[5] => -0.1095657
)
[2] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => 0.5250869
)
[3] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.0686326
)
[4] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => 0.4661818
)
[5] => Array
(
[0] => 1
[1] => 1
[2] => 15.99491464
[3] => 3.654
[4] => 3.654
[5] => -0.9945602
)
[6] => Array
(
[0] => 1
[1] => 1
[2] => 30.9737634
[3] => 2.515
[4] => 2.515
[5] => 1.883921
)
[7] => Array
(
[0] => 1
[1] => 1
[2] => 15.99491464
[3] => 3.654
[4] => 3.654
[5] => -1.095338
)
[8] => Array
(
[0] => 1
[1] => 1
[2] => 15.99491464
[3] => 3.654
[4] => 3.654
[5] => -1.019472
)
[9] => Array
(
[0] => 1
[1] => 4
[2] => 12
[3] => 2.746
[4] => 2.6296707827796
[5] => 0.4202826
)
)
[e] => seed 0
[f] => forcefield mm+
[m] => mol 1
[s] => sys 0 0 1
)
Fragments tree structure: Array
(
[1] => Array
(
[2] => Array
(
[0] => 1
)
[3] => Array
(
[0] => 1
)
[4] => Array
(
[0] => 1
[1] => 2
[2] => 7
)
[5] => Array
(
[0] => 1
[1] => 2
)
[6] => Array
(
[0] => 1
[1] => 2
[2] => 3
)
[7] => Array
(
[0] => 1
)
[8] => Array
(
[0] => 1
[1] => 2
[2] => 3
)
[9] => Array
(
[0] => 1
[1] => 2
[2] => 3
[3] => 4
[4] => 7
)
[1] => Array
(
[0] => 0
)
)
[2] => Array
(
[1] => Array
(
[0] => 2
[1] => 3
[2] => 4
[3] => 5
[4] => 6
[5] => 7
[6] => 8
[7] => 9
)
[3] => Array
(
[0] => 2
[1] => 1
[2] => 6
[3] => 7
[4] => 8
[5] => 9
)
[4] => Array
(
[0] => 2
[1] => 1
[2] => 7
)
[5] => Array
(
[0] => 2
[1] => 1
[2] => 3
[3] => 7
)
[6] => Array
(
[0] => 2
[1] => 1
[2] => 3
)
[7] => Array
(
[0] => 2
[1] => 1
[2] => 3
[3] => 4
)
[8] => Array
(
[0] => 2
[1] => 1
[2] => 3
[3] => 4
[4] => 7
)
[9] => Array
(
[0] => 2
[1] => 1
[2] => 3
[3] => 4
[4] => 7
)
[2] => Array
(
[0] => 0
)
)
[3] => Array
(
[1] => Array
(
[0] => 3
[1] => 4
[2] => 5
)
[2] => Array
(
[0] => 3
[1] => 4
[2] => 5
)
[4] => Array
(
[0] => 3
[1] => 1
[2] => 2
[3] => 7
)
[5] => Array
(
[0] => 3
[1] => 1
[2] => 2
)
[6] => Array
(
[0] => 3
[1] => 1
[2] => 2
[3] => 4
)
[7] => Array
(
[0] => 3
[1] => 4
)
[8] => Array
(
[0] => 3
[1] => 1
[2] => 2
[3] => 4
)
[9] => Array
(
[0] => 3
[1] => 1
[2] => 2
[3] => 4
[4] => 5
[5] => 7
)
[3] => Array
(
[0] => 0
)
)
[4] => Array
(
[1] => Array
(
[0] => 4
[1] => 3
[2] => 5
[3] => 6
[4] => 8
[5] => 9
)
[2] => Array
(
[0] => 4
[1] => 5
)
[3] => Array
(
[0] => 4
[1] => 5
[2] => 6
[3] => 8
[4] => 9
)
[5] => Array
(
[0] => 4
[1] => 1
[2] => 2
[3] => 3
)
[6] => Array
(
[0] => 4
[1] => 3
)
[7] => Array
(
[0] => 4
[1] => 3
[2] => 5
)
[8] => Array
(
[0] => 4
[1] => 1
[2] => 2
[3] => 3
[4] => 5
)
[9] => Array
(
[0] => 4
[1] => 1
[2] => 2
[3] => 3
[4] => 5
)
[4] => Array
(
[0] => 0
)
)
[5] => Array
(
[1] => Array
(
[0] => 5
[1] => 4
[2] => 6
[3] => 8
[4] => 9
)
[2] => Array
(
[0] => 5
[1] => 4
[2] => 6
[3] => 8
[4] => 9
)
[3] => Array
(
[0] => 5
[1] => 6
[2] => 8
[3] => 9
)
[4] => Array
(
[0] => 5
[1] => 6
[2] => 7
[3] => 8
[4] => 9
)
[6] => Array
(
[0] => 5
[1] => 3
[2] => 4
)
[7] => Array
(
[0] => 5
[1] => 4
)
[8] => Array
(
[0] => 5
[1] => 3
[2] => 4
)
[9] => Array
(
[0] => 5
[1] => 1
[2] => 2
[3] => 3
[4] => 4
[5] => 6
[6] => 7
)
[5] => Array
(
[0] => 0
)
)
[6] => Array
(
[1] => Array
(
[0] => 6
[1] => 4
[2] => 5
[3] => 7
[4] => 8
[5] => 9
)
[2] => Array
(
[0] => 6
[1] => 5
[2] => 8
[3] => 9
)
[3] => Array
(
[0] => 6
[1] => 5
[2] => 7
[3] => 8
[4] => 9
)
[4] => Array
(
[0] => 6
[1] => 7
[2] => 8
[3] => 9
)
[5] => Array
(
[0] => 6
[1] => 1
[2] => 2
[3] => 7
[4] => 8
[5] => 9
)
[7] => Array
(
[0] => 6
[1] => 4
[2] => 5
[3] => 8
[4] => 9
)
[8] => Array
(
[0] => 6
[1] => 1
[2] => 2
[3] => 3
[4] => 4
[5] => 5
[6] => 7
)
[9] => Array
(
[0] => 6
[1] => 1
[2] => 2
[3] => 3
[4] => 4
[5] => 5
[6] => 7
)
[6] => Array
(
[0] => 0
)
)
[7] => Array
(
[1] => Array
(
[0] => 7
[1] => 5
[2] => 6
[3] => 8
[4] => 9
)
[2] => Array
(
[0] => 7
[1] => 5
[2] => 6
[3] => 8
[4] => 9
)
[3] => Array
(
[0] => 7
[1] => 6
[2] => 8
[3] => 9
)
[4] => Array
(
[0] => 7
[1] => 1
[2] => 2
[3] => 6
[4] => 8
[5] => 9
)
[5] => Array
(
[0] => 7
[1] => 1
[2] => 2
)
[6] => Array
(
[0] => 7
[1] => 1
[2] => 2
[3] => 3
)
[8] => Array
(
[0] => 7
[1] => 1
[2] => 2
[3] => 3
)
[9] => Array
(
[0] => 7
[1] => 1
[2] => 2
[3] => 3
[4] => 4
[5] => 5
[6] => 6
)
[7] => Array
(
[0] => 0
)
)
[8] => Array
(
[1] => Array
(
[0] => 8
[1] => 5
[2] => 6
[3] => 9
)
[2] => Array
(
[0] => 8
[1] => 5
[2] => 6
[3] => 9
)
[3] => Array
(
[0] => 8
[1] => 6
[2] => 9
)
[4] => Array
(
[0] => 8
[1] => 6
[2] => 7
[3] => 9
)
[5] => Array
(
[0] => 8
[1] => 9
)
[6] => Array
(
[0] => 8
[1] => 9
)
[7] => Array
(
[0] => 8
[1] => 9
)
[9] => Array
(
[0] => 8
[1] => 1
[2] => 2
[3] => 3
[4] => 4
[5] => 5
[6] => 6
[7] => 7
)
[8] => Array
(
[0] => 0
)
)
[9] => Array
(
[1] => Array
(
[0] => 9
[1] => 5
[2] => 6
[3] => 8
)
[2] => Array
(
[0] => 9
[1] => 8
)
[3] => Array
(
[0] => 9
[1] => 6
[2] => 8
)
[4] => Array
(
[0] => 9
[1] => 8
)
[5] => Array
(
[0] => 9
[1] => 8
)
[6] => Array
(
[0] => 9
)
[7] => Array
(
[0] => 9
[1] => 8
)
[8] => Array
(
[0] => 9
)
[9] => Array
(
[0] => 0
)
)
)
Fragments: 72
Fragmental properties: Array
(
[0] => 144
[1] => 18
[2] => 32.808788679292
[3] => 7.5833333333333
[4] => 28.666666666667
[5] => 18
[6] => 9.8333333333333
[7] => 16.279455345959
[8] => 243.99228165407
[9] => 234.66149059666
[10] => 32.808788679292
[11] => 57.562995125944
[12] => 28.666666666667
[13] => 185.33333333333
[14] => 47.146328459277
[15] => 16.279455345959
[16] => 27.33079105741
[17] => 171.47545534596
[18] => 176.80878867929
[19] => 25.583333333333
[20] => 46.666666666667
[21] => 23.333333333333
[22] => 15.166666666667
[23] => 25.754910691918
[24] => 68.012619861456
[25] => 27.475455345959
[26] => 216.66149059666
[27] => 169.58333333333
[28] => 23.333333333333
[29] => 55.312995125944
[30] => 47.146328459277
[31] => 16.279455345959
[32] => 50.012619861456
[33] => 159.12266308681
[34] => 72.661490596657
[35] => 396.9599574277
[36] => 279.17062767422
[37] => 37.312995125944
[38] => 39.562995125944
[39] => 145.67666308681
[40] => 81.992281654066
[41] => 141.12266308681
[42] => 104.64115238927
[43] => 141.12266308681
[44] => 1036.2500094111
[45] => 1034.0000094111
[46] => 1038.5000094111
[47] => 145.67666308681
[48] => 76.658948320733
[49] => 396.9599574277
[50] => 72.661490596657
[51] => 164.45599642015
[52] => 39.562995125944
[53] => 284.50396100755
[54] => 41.812995125944
[55] => 145.67666308681
[56] => 44.679286528122
[57] => 141.12266308681
[58] => 40.681828804046
[59] => 141.12266308681
[60] => 37.312995125944
[61] => 273.83729434089
[62] => 37.312995125944
[63] => 401.5139574277
[64] => 44.679286528122
[65] => 11.725455345959
[66] => 40.681828804046
[67] => 11.725455345959
[68] => 37.312995125944
[69] => 18
[70] => 37.312995125944
[71] => 144
)
GDmSMt = 67.772137360531
lGDmSMt = 4.2161511570974
Demonstrative calculations for mr10 molecule and lAmrfEt MDF member
POST data: Array
(
[hin] => 10_mr1010.hin
[Do] => t
[Ap] => E
[Df] => f
[Im] => r
[Fc] => m
[Sf] => A
[Lo] => l
)
Molecule's data: m_c Object
(
[a] => 12
[b] => 13
[atom] => Array
(
[1] => Array
(
[0] => 1
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.01363325
[6] => -1.374815
[7] => -2.525667
[8] => -1.163645
[9] => 2
[10] => 2
[11] => 1
[12] => 6
[13] => 1
)
[2] => Array
(
[0] => 2
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.03494024
[6] => -0.6541022
[7] => -3.076405
[8] => 0.08466685
[9] => 2
[10] => 1
[11] => 1
[12] => 3
[13] => 1
)
[3] => Array
(
[0] => 3
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.01286221
[6] => 0.8641648
[7] => -2.810718
[8] => 0.0138855
[9] => 2
[10] => 2
[11] => 1
[12] => 4
[13] => 1
)
[4] => Array
(
[0] => 4
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.2916856
[6] => 1.165918
[7] => -1.313278
[8] => 0.1745337
[9] => 2
[10] => 3
[11] => 1
[12] => 5
[13] => 1
)
[5] => Array
(
[0] => 5
[1] => -
[2] => P
[3] => P5
[4] => -
[5] => 0.9998846
[6] => 0.3477604
[7] => -0.3762538
[8] => -1.163645
[9] => 3
[10] => 4
[11] => 1
[12] => 6
[13] => 1
[14] => 7
[15] => 1
)
[6] => Array
(
[0] => 6
[1] => -
[2] => C
[3] => C4
[4] => -
[5] => -0.07605362
[6] => -1.374815
[7] => -0.9852762
[8] => -1.163645
[9] => 2
[10] => 5
[11] => 1
[12] => 1
[13] => 1
)
[7] => Array
(
[0] => 7
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.2473259
[6] => 0.2043564
[7] => 1.029266
[8] => -0.0003523827
[9] => 3
[10] => 5
[11] => 1
[12] => 8
[13] => 2
[14] => 9
[15] => 1
)
[8] => Array
(
[0] => 8
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.04595566
[6] => -0.4518933
[7] => 0.876143
[8] => 1.157674
[9] => 2
[10] => 7
[11] => 2
[12] => 10
[13] => 1
)
[9] => Array
(
[0] => 9
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.04757643
[6] => 0.7997317
[7] => 2.319123
[8] => -0.3369715
[9] => 2
[10] => 7
[11] => 1
[12] => 11
[13] => 2
)
[10] => Array
(
[0] => 10
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.04881668
[6] => -0.5716058
[7] => 1.999108
[8] => 2.082884
[9] => 2
[10] => 8
[11] => 1
[12] => 12
[13] => 2
)
[11] => Array
(
[0] => 11
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.0406456
[6] => 0.6907468
[7] => 3.349581
[8] => 0.5127543
[9] => 2
[10] => 9
[11] => 2
[12] => 12
[13] => 1
)
[12] => Array
(
[0] => 12
[1] => -
[2] => C
[3] => C3
[4] => -
[5] => -0.05546284
[6] => -0.02339876
[7] => 3.182384
[8] => 1.775193
[9] => 2
[10] => 10
[11] => 2
[12] => 11
[13] => 1
)
)
[prop] => Array
(
[1] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.01363325
)
[2] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.03494024
)
[3] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.01286221
)
[4] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.2916856
)
[5] => Array
(
[0] => 1
[1] => 1
[2] => 30.9737634
[3] => 2.515
[4] => 2.515
[5] => 0.9998846
)
[6] => Array
(
[0] => 1
[1] => 3
[2] => 12
[3] => 2.746
[4] => 2.6423489837131
[5] => -0.07605362
)
[7] => Array
(
[0] => 1
[1] => 1
[2] => 12
[3] => 2.746
[4] => 2.746
[5] => -0.2473259
)
[8] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => -0.04595566
)
[9] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => -0.04757643
)
[10] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => -0.04881668
)
[11] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => -0.0406456
)
[12] => Array
(
[0] => 1
[1] => 2
[2] => 12
[3] => 2.746
[4] => 2.6678890531654
[5] => -0.05546284
)
)
[e] => seed 0
[f] => forcefield mm+
[m] => mol 1
[s] => sys 0 0 1
)
Fragments tree structure: Array
(
[1] => Array
(
[1] => Array
(
[0] => 0
)
[2] => Array
(
[0] => 1
)
[3] => Array
(
[0] => 1
)
[4] => Array
(
[0] => 1
)
[5] => Array
(
[0] => 1
)
[6] => Array
(
[0] => 1
)
[7] => Array
(
[0] => 1
)
[8] => Array
(
[0] => 1
)
[9] => Array
(
[0] => 1
)
[10] => Array
(
[0] => 1
)
[11] => Array
(
[0] => 1
)
[12] => Array
(
[0] => 1
)
)
[2] => Array
(
[1] => Array
(
[0] => 2
)
[2] => Array
(
[0] => 0
)
[3] => Array
(
[0] => 2
)
[4] => Array
(
[0] => 2
)
[5] => Array
(
[0] => 2
)
[6] => Array
(
[0] => 2
)
[7] => Array
(
[0] => 2
)
[8] => Array
(
[0] => 2
)
[9] => Array
(
[0] => 2
)
[10] => Array
(
[0] => 2
)
[11] => Array
(
[0] => 2
)
[12] => Array
(
[0] => 2
)
)
[3] => Array
(
[1] => Array
(
[0] => 3
)
[2] => Array
(
[0] => 3
)
[3] => Array
(
[0] => 0
)
[4] => Array
(
[0] => 3
)
[5] => Array
(
[0] => 3
)
[6] => Array
(
[0] => 3
)
[7] => Array
(
[0] => 3
)
[8] => Array
(
[0] => 3
)
[9] => Array
(
[0] => 3
)
[10] => Array
(
[0] => 3
)
[11] => Array
(
[0] => 3
)
[12] => Array
(
[0] => 3
)
)
[4] => Array
(
[1] => Array
(
[0] => 4
)
[2] => Array
(
[0] => 4
)
[3] => Array
(
[0] => 4
)
[4] => Array
(
[0] => 0
)
[5] => Array
(
[0] => 4
)
[6] => Array
(
[0] => 4
)
[7] => Array
(
[0] => 4
)
[8] => Array
(
[0] => 4
)
[9] => Array
(
[0] => 4
)
[10] => Array
(
[0] => 4
)
[11] => Array
(
[0] => 4
)
[12] => Array
(
[0] => 4
)
)
[5] => Array
(
[1] => Array
(
[0] => 5
)
[2] => Array
(
[0] => 5
)
[3] => Array
(
[0] => 5
)
[4] => Array
(
[0] => 5
)
[5] => Array
(
[0] => 0
)
[6] => Array
(
[0] => 5
)
[7] => Array
(
[0] => 5
)
[8] => Array
(
[0] => 5
)
[9] => Array
(
[0] => 5
)
[10] => Array
(
[0] => 5
)
[11] => Array
(
[0] => 5
)
[12] => Array
(
[0] => 5
)
)
[6] => Array
(
[1] => Array
(
[0] => 6
)
[2] => Array
(
[0] => 6
)
[3] => Array
(
[0] => 6
)
[4] => Array
(
[0] => 6
)
[5] => Array
(
[0] => 6
)
[6] => Array
(
[0] => 0
)
[7] => Array
(
[0] => 6
)
[8] => Array
(
[0] => 6
)
[9] => Array
(
[0] => 6
)
[10] => Array
(
[0] => 6
)
[11] => Array
(
[0] => 6
)
[12] => Array
(
[0] => 6
)
)
[7] => Array
(
[1] => Array
(
[0] => 7
)
[2] => Array
(
[0] => 7
)
[3] => Array
(
[0] => 7
)
[4] => Array
(
[0] => 7
)
[5] => Array
(
[0] => 7
)
[6] => Array
(
[0] => 7
)
[7] => Array
(
[0] => 0
)
[8] => Array
(
[0] => 7
)
[9] => Array
(
[0] => 7
)
[10] => Array
(
[0] => 7
)
[11] => Array
(
[0] => 7
)
[12] => Array
(
[0] => 7
)
)
[8] => Array
(
[1] => Array
(
[0] => 8
)
[2] => Array
(
[0] => 8
)
[3] => Array
(
[0] => 8
)
[4] => Array
(
[0] => 8
)
[5] => Array
(
[0] => 8
)
[6] => Array
(
[0] => 8
)
[7] => Array
(
[0] => 8
)
[8] => Array
(
[0] => 0
)
[9] => Array
(
[0] => 8
)
[10] => Array
(
[0] => 8
)
[11] => Array
(
[0] => 8
)
[12] => Array
(
[0] => 8
)
)
[9] => Array
(
[1] => Array
(
[0] => 9
)
[2] => Array
(
[0] => 9
)
[3] => Array
(
[0] => 9
)
[4] => Array
(
[0] => 9
)
[5] => Array
(
[0] => 9
)
[6] => Array
(
[0] => 9
)
[7] => Array
(
[0] => 9
)
[8] => Array
(
[0] => 9
)
[9] => Array
(
[0] => 0
)
[10] => Array
(
[0] => 9
)
[11] => Array
(
[0] => 9
)
[12] => Array
(
[0] => 9
)
)
[10] => Array
(
[1] => Array
(
[0] => 10
)
[2] => Array
(
[0] => 10
)
[3] => Array
(
[0] => 10
)
[4] => Array
(
[0] => 10
)
[5] => Array
(
[0] => 10
)
[6] => Array
(
[0] => 10
)
[7] => Array
(
[0] => 10
)
[8] => Array
(
[0] => 10
)
[9] => Array
(
[0] => 10
)
[10] => Array
(
[0] => 0
)
[11] => Array
(
[0] => 10
)
[12] => Array
(
[0] => 10
)
)
[11] => Array
(
[1] => Array
(
[0] => 11
)
[2] => Array
(
[0] => 11
)
[3] => Array
(
[0] => 11
)
[4] => Array
(
[0] => 11
)
[5] => Array
(
[0] => 11
)
[6] => Array
(
[0] => 11
)
[7] => Array
(
[0] => 11
)
[8] => Array
(
[0] => 11
)
[9] => Array
(
[0] => 11
)
[10] => Array
(
[0] => 11
)
[11] => Array
(
[0] => 0
)
[12] => Array
(
[0] => 11
)
)
[12] => Array
(
[1] => Array
(
[0] => 12
)
[2] => Array
(
[0] => 12
)
[3] => Array
(
[0] => 12
)
[4] => Array
(
[0] => 12
)
[5] => Array
(
[0] => 12
)
[6] => Array
(
[0] => 12
)
[7] => Array
(
[0] => 12
)
[8] => Array
(
[0] => 12
)
[9] => Array
(
[0] => 12
)
[10] => Array
(
[0] => 12
)
[11] => Array
(
[0] => 12
)
[12] => Array
(
[0] => 0
)
)
)
Fragments: 132
Fragmental properties: Array
(
[0] => 7.540516
[1] => 1.885129
[2] => 0.83783511111111
[3] => 1.7265475
[4] => 7.540516
[5] => 0.83783511111111
[6] => 0.47128225
[7] => 0.47128225
[8] => 0.30162064
[9] => 0.30162064
[10] => 0.20945877777778
[11] => 7.540516
[12] => 7.540516
[13] => 1.885129
[14] => 0.76735444444444
[15] => 1.885129
[16] => 0.47128225
[17] => 0.30162064
[18] => 0.30162064
[19] => 0.20945877777778
[20] => 0.20945877777778
[21] => 0.15388808163265
[22] => 1.885129
[23] => 7.540516
[24] => 7.540516
[25] => 1.7265475
[26] => 0.83783511111111
[27] => 0.83783511111111
[28] => 0.47128225
[29] => 0.47128225
[30] => 0.30162064
[31] => 0.30162064
[32] => 0.20945877777778
[33] => 0.83783511111111
[34] => 1.885129
[35] => 7.540516
[36] => 6.90619
[37] => 1.885129
[38] => 1.885129
[39] => 0.83783511111111
[40] => 0.83783511111111
[41] => 0.47128225
[42] => 0.47128225
[43] => 0.30162064
[44] => 1.7265475
[45] => 0.76735444444444
[46] => 1.7265475
[47] => 6.90619
[48] => 6.90619
[49] => 6.90619
[50] => 1.7265475
[51] => 1.7265475
[52] => 0.76735444444444
[53] => 0.76735444444444
[54] => 0.431636875
[55] => 7.540516
[56] => 1.885129
[57] => 0.83783511111111
[58] => 1.885129
[59] => 6.90619
[60] => 1.885129
[61] => 0.83783511111111
[62] => 0.83783511111111
[63] => 0.47128225
[64] => 0.47128225
[65] => 0.30162064
[66] => 0.83783511111111
[67] => 0.47128225
[68] => 0.83783511111111
[69] => 1.885129
[70] => 6.90619
[71] => 1.885129
[72] => 7.540516
[73] => 7.540516
[74] => 1.885129
[75] => 1.885129
[76] => 0.83783511111111
[77] => 0.47128225
[78] => 0.30162064
[79] => 0.47128225
[80] => 0.83783511111111
[81] => 1.7265475
[82] => 0.83783511111111
[83] => 7.540516
[84] => 1.885129
[85] => 7.540516
[86] => 0.83783511111111
[87] => 1.885129
[88] => 0.47128225
[89] => 0.30162064
[90] => 0.47128225
[91] => 0.83783511111111
[92] => 1.7265475
[93] => 0.83783511111111
[94] => 7.540516
[95] => 1.885129
[96] => 0.83783511111111
[97] => 7.540516
[98] => 1.885129
[99] => 0.30162064
[100] => 0.20945877777778
[101] => 0.30162064
[102] => 0.47128225
[103] => 0.76735444444444
[104] => 0.47128225
[105] => 1.885129
[106] => 7.540516
[107] => 0.83783511111111
[108] => 1.885129
[109] => 7.540516
[110] => 0.30162064
[111] => 0.20945877777778
[112] => 0.30162064
[113] => 0.47128225
[114] => 0.76735444444444
[115] => 0.47128225
[116] => 1.885129
[117] => 0.83783511111111
[118] => 7.540516
[119] => 1.885129
[120] => 7.540516
[121] => 0.20945877777778
[122] => 0.15388808163265
[123] => 0.20945877777778
[124] => 0.30162064
[125] => 0.431636875
[126] => 0.30162064
[127] => 0.83783511111111
[128] => 1.885129
[129] => 1.885129
[130] => 7.540516
[131] => 7.540516
)
AmrfEt = 2.2007209544439
lAmrfEt = 0.78878501324562